A 100-fold leap to GigaDalton DNA nanotech

DNA, present in almost every cell, is increasingly being used as a building material to construct tiny, but sophisticated structures such as autonomous 'DNA walkers' that can move along a microparticle surface, fluorescent labels for diagnostic applications, 'DNA boxes' that serve as smart drug-delivery vehicles programmed to open up at disease sites to release their therapeutic content, or programmable factories for nanoparticles of defined sizes and shapes for new optical and electronic applications.
To accommodate these functions, researchers at Harvard's Wyss Institute for Biologically Inspired Engineering and around the world have developed ways that allow DNA strands to self-assemble into increasingly complex 3D structures such as scaffolded DNA origamis. DNA origamis, however, are limited in their sizes because they rely on the availability of scaffold strands that can be difficult to manufacture and manipulate. In 2012, Peng Yin and his team at the Wyss Institute presented an alternative method in Nature (2D) and Science (3D) that is based on DNA 'bricks', which do not use a scaffold but rather are able to connect like interlocking Lego bricks and thereby self-assemble into origami-sized structures with prescribed shapes.
As reported in Nature, the team leapfrogged their technology by two orders of magnitude, enabling next-generation DNA bricks to self-assemble into three-dimensional nanostructures that are 100 times more complex than those created with existing methods. DNA origami and first generation DNA bricks self-assemble from hundreds of unique components to produce nanostructures on the MegaDalton scale, whereas the new DNA bricks approach allows 10,000 components to self-assemble into GigaDalton-sized structures (1 GigaDalton equals 1000 MegaDaltons or 1 billion Daltons). The study provides user-friendly computational tools to design DNA nanostructures with complex cavities (and possibly surfaces) that have the potential to serve as building components in numerous nanotechnological applications in medicine and engineering.
"The principle and promising capabilities of our first-generation DNA bricks led us to ask whether we can enhance the system to attain significantly more complex nanostructures with much higher yields in one-pot assembly reactions. Here we managed to do all this. We worked out an easily accessible practical platform that allows researchers with very different interests and applications in mind to create a molecular canvas with 10,000 bricks and use it to build nanostructures with unprecedented complexities and potential," said corresponding author Yin, Ph.D., who is a Wyss Institute Core Faculty member, co-leader of the Institute's Molecular Robotics Initiative, and Professor of Systems Biology at Harvard Medical School.
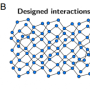
DNA brick technology is based on the stable and highly programmable nature of DNA. A single DNA brick is a short strand of synthetic DNA made up of a pre-defined sequence of the four universal nucleotide bases: adenine (A), cytosine (C), guanine (G), and thymine (T). The Wyss Institute's researchers create large 3D nanostructures by mixing various bricks, each carrying its own unique sequence of nucleotides that is designed to fit and bind to a complimentary domain of nucleotide bases in another brick so that they can self-assemble. In the technology's new version, by varying the length of individual binding domains within the bricks, the team ended up with a substantially increased diversity among possible bricks that, in addition, bind much stronger to each other. The study also developed a user-friendly computer software so designers can simply input a required 3D shape and automatically receive a list of DNA brick sequences that can be synthesized and used to form the desired structure.
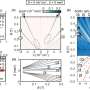
"We demonstrated the capabilities of our technology by constructing massive cuboids containing up to 30,000 bricks and showed a few exemplary shapes that can be built from subsets of those bricks. It is remarkable that the bricks were able to distinguish between tens of thousands of potential partners to find their correct neighbors, and it was exciting to see that the DNA bricks technique could be used to form rather complex cavities such as a teddy bear, the word 'LOVE' or a Möbius strip, amongst many others, " said first author Luvena Ong, Ph.D., a former Graduate Student in Yin's laboratory and now a Research Investigator at Bristol-Myers Squibb.
Yin's team collaborated with researchers at the National Center for Scientific Research (CNRS) and the French National Institute of Health and Medical Research (INSERM) in Montpellier, France and the Max Planck Institute of Biochemistry in Munich, Germany to deploy a collection of state-of-the-art microscopy methods to visualize the designed cavities in 3D cuboids. "Cavity structures composed of DNA bricks are of much interest as they offer the possibility to design nano-containers in which biomolecules like proteins can be place in very defined arrangements to study their interactions and leverage their activities," said co-corresponding author Yonggang Ke, Ph.D., who developed the first DNA brick platform with Yin as a Postdoctoral Fellow at the Wyss Institute, and is now Assistant Professor at the Georgia Institute of Technology and Emory University. Ke, working together with his Graduate Student Pengfei Wang, was instrumental in advancing the technology to its new version. "By adding functional moieties to DNA bricks that can carry out assembly and enzymatic processes, they can be converted into powerful tools for commercial and biomedical nanofabrication processes on a new scale," said Ke. The researchers believe that, in the future, the method could also be used to generate large nanostructures with sculpted and application-specific outer surfaces.
"The way the multifaceted DNA bricks technology is evolving shows how the Wyss Institute's Molecular Robotics Initiative can reach deep into the field of DNA nanotechnology to enable new approaches that could solve many real world problems," said Wyss Institute Founding Director Donald Ingber, M.D., Ph.D., who is also the Judah Folkman Professor of Vascular Biology at HMS and the Vascular Biology Program at Boston Children's Hospital, as well as Professor of Bioengineering at SEAS.
More information: Gigadalton-scale shape-programmable DNA assemblies, Nature (2017). nature.com/articles/doi:10.1038/nature24651
Provided by Harvard University