Improving nature's tools for digesting plastic

Enzymes found in nature can break down certain plastics, but not well enough to support industrial recycling and stem the scourge of plastic waste. Building on what nature has provided, researchers at Rensselaer Polytechnic Institute have improved the efficiency of a leaf and branch compost cutinase that breaks down polyethylene terephthalate (PET), the plastic used in clear and colored plastic water bottles and many other products. Researchers believe the enzyme can be further refined, offering a promising candidate to fuel limitless recycling of PET and possibly other plastics such as cellulose acetate.
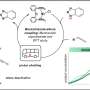
In work recently published in the journal Biochemistry, the researchers used yeast cells to express the leaf and branch compost cutinase (LCC) modified by the addition of sugar molecules – or glycans – in two locations. The "glycosylated" modified enzyme retained at least half of its activity after 48 hours at 75 degrees Celsius, versus a previously reported half-life of 40 minutes for the unmodified enzyme at 70 degrees Celsius.
"We need plastics and other materials that retain good performance and, after use, can then be broken down by safe and mild processes to their original building blocks for reuse," said Richard Gross, lead author of the research, Constellation Professor of Biocatalysis and Metabolic Engineering, member of the Center for Biotechnology and Interdisciplinary Studies, and Professor of Chemistry and Chemical Biology at Rensselaer. "The goal should be zero waste and to do that, we have to build reuse into the design of a wide range of polymers and materials. This is an encouraging step toward that goal."
"This promising advance, which is sorely needed as plastic pollution becomes an ever-greater threat to our environment, is a result of the diverse skill set and collaborative environment we have built at Rensselaer " said Deepak Vashishth, director of the Center for Biotechnology and Interdisciplinary Studies. "Dr. Gross' research spans boundaries between biologics and biomanufacturing, and is certain to help us resolve the critical problems we face."
With existing technologies, a plastic bottle isn't so much recycled as down-cycled. After a single use, a high percentage of PET bottles go directly to landfills or are reused as other plastics such as PET fibers and fleece for clothes, carpet, bags, furniture, and packing materials. Eventually, down-cycled PET makes its way to landfills or other undesirable environments such as oceans and lakes, a fate many consumers are unaware of as they toss their water bottles in a recycling bin.
Breaking PET down into its building blocks – terephthalic acid and ethylene glycol – would enable the limitless reuse more commonly associated with other recyclable materials such as glass and metal. Some naturally occurring enzymes can break down PET, but not within the constraints of time and temperature required by an industrial recycling process. Many enzymes lose their activity at higher temperatures, and eventually denature. An enzyme suitable for industrial recycling must be able to operate at optimal temperature for breaking down PET, which is about 75 degree Celsius, and it must retain its activity long enough to do its job cost-effectively at that temperature.
LCC was initially discovered through metagenomic analysis of a leaf-branch compost, meaning scientists extracted DNA found in a compost irrespective of the organisms that produced it, and then used the DNA to express and catalogue enzymes that were present. A 2012 study published by unrelated researchers in the journal Applied and Environmental Microbiology showed that LCC was able to hydrolyze, or break down, PET, but lost activity quickly at higher temperatures. That attracted the attention of Gross, an expert on biocatalytic and chemical synthetic methods, who saw the opportunity to improve the enzyme's "kinetic stability" without damaging its ability to break down PET.
The lab studied the enzyme and found three separate glycosylation sites, amino acid sequences to which glycans are attached during protein synthesis. Gross said the glycosylation sites could have evolved in a previous organism and been conserved even though they were not used by the natural bacterium that originally produced this protein. Regardless, when the team expressed the enzyme using the yeast strain Pichia pastoris, they found that the yeast naturally glycosylated the enzyme at the three sites. Further research showed that two glycosylation sites yielded a more effective enzyme than three sites.
With only those minor changes, the team saw greater than a 60-fold improvement in kinetic stability. And Gross said additional research will explore how to further improve the kinetics and the overall activity of the enzyme by experimenting with amino acid sequences to create variant structures. Through this work, Gross expects to understand the design rules that lead to better performance.
"This cutinase is an excellent candidate for commercialization, but this work will also help us redesign other cutinases to break down other polymers, and that's a much larger end game," said Gross.
"Stabilizing Leaf and Branch Compost Cutinase (LCC) with Glycosylation: Mechanism and Effect on PET Hydrolysis" was published in Biochemistry.
More information: Abhijit N. Shirke et al. Stabilizing Leaf and Branch Compost Cutinase (LCC) with Glycosylation: Mechanism and Effect on PET Hydrolysis, Biochemistry (2018). DOI: 10.1021/acs.biochem.7b01189
Journal information: Biochemistry , Applied and Environmental Microbiology
Provided by Rensselaer Polytechnic Institute